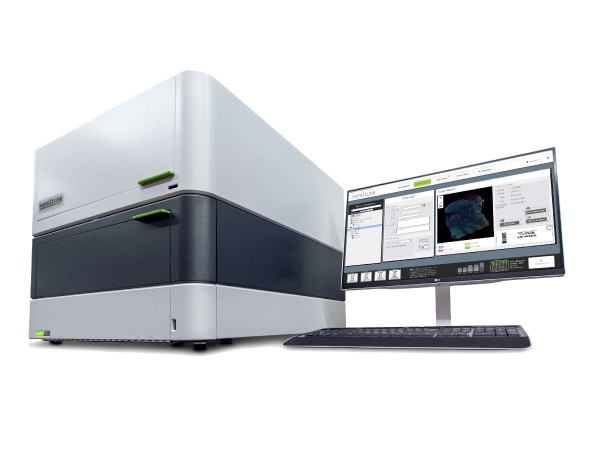
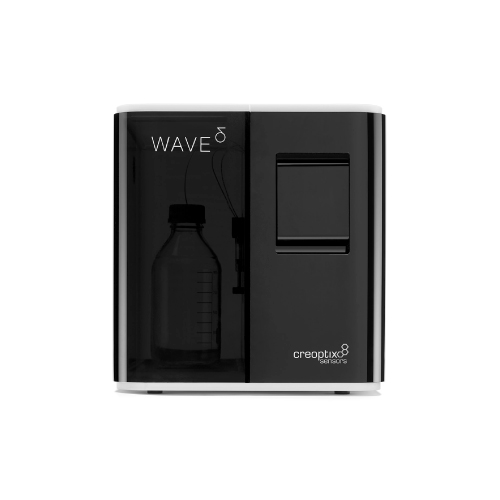
Label-free, Real-time, Molecular Interaction analysis system :WAVEsystem®

feature
- 1
GCI (Grating-coupled Interferometry) 원리 및 Label-free 방식으로 실시간 결합 반응 측정
Creoptix사의 Creoptix ® WAVEsystem®은 라벨 프리 샘플의 실시간 물질 결합반응 분석 (친화도, 카이네틱스) 측정을 고감도, 고속으로 실시할 수 있는 물질 상호작용 분자 시스템입니다. 독자적인 Grating-coupled Interferometry (GCI, 도파관 간섭법) 기술과 견고한 Microfluidics engineering을 조합해 종래 기술에서는 측정이 곤란했던 샘플을 높은 감도, 고속 측정을 실현했습니다.
- 2
분석 가능 물질 (친화도, 카이네틱스)
Crude Sample (혈청 또는 혈장, 항체, 바이러스 유사 입자, 바이러스, 리포좀, 막 단백질), 소분자/단편 라이브러리, 큰 분자 등 다양한 검체에 대응하여 연구를 가속화시킬 수 있습니다.
- 3
고감도 측정(친화도, 카이네틱스)
Rmax 1 pg/mm2 이하의 신호도 신뢰할 수 있는 고감도
Signal to noise 0.01 pg/mm2 이하의 신호까지 신뢰 가능한 고감도
분자량비 1000:1을 넘는 작용기 단백질 (ligand)와 약물 (analyte)를 분석 가능 - 4
고속, 대량 측정
waveRAPID® 분석방법으로 분석 시간을 대폭 단축시키고 high throughput testing 실현
waveRAPID® 분석방법은 1회 주입만으로도 결합 반응 (카이네틱스) 측정이 가능한 방법입니다.
자세한 내용은 아래 내용을 참고 부탁드립니다.
* 예: 90 여개의 저분자 화합물 분석을 18 시간내 분석 가능 (기존 multi-cycle kinetics의 경우, 6배 이상의 시간 소요됨)
※ waveRAPID®는 WAVEdelta 모델에서만 작동합니다. 외관은 같으며, 자세한 내용은 specification란을 확인 부탁드립니다.
- 5
샘플당 칩 활용 단가가 낮고 사용이 편리한 WAVEchip
- 교차오염이 없고 쉬운 유지 보수
- 마이크로 밸브 없는 디자인
- 아세토니트릴, 고농도의 DMSO 등의 용매에 대응
- 폭넓은 샘플에 적응하는 풍부한 라인업
- Crude sample (혈청 또는 혈장, 항체, 바이러스-유사 입자, 바이러스, 리포좀, 막 단백질), 저분자/단편 라이브러리, 난이도가 높은 화합물, 1000nm까지의 입자에 대응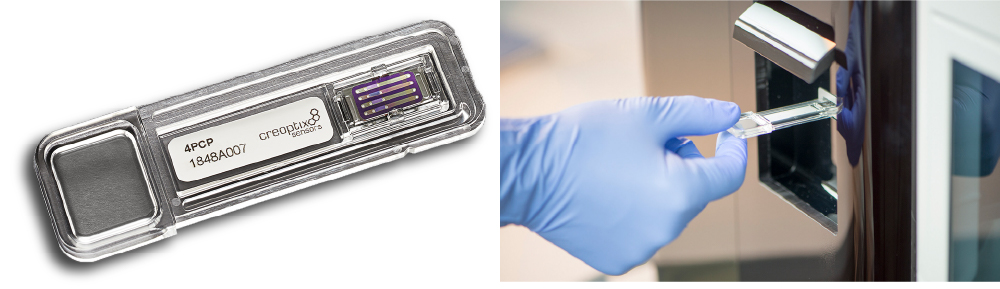
- 6
폭넓은 측정 범위: koff =10-5 ~10 s-1
카트리지 설계에 의한 150ms (밀리초)의 초고속 전이 시간을 실현, 빠른 off-rate 분해능을 제공(KD (Equilibrium constant)의 범위는 1 pM ~ 1 mM 입니다.)
- 7
직관적인 조작으로 쉽게 실험 수행 및 결과 해석이 가능한 WAVEcontrol 소프트웨어
- 8
Autosampler는 48 vial rack, 96/384 well plate 두개를 채용하여 최대 1~768 개 샘플 탑재 가능
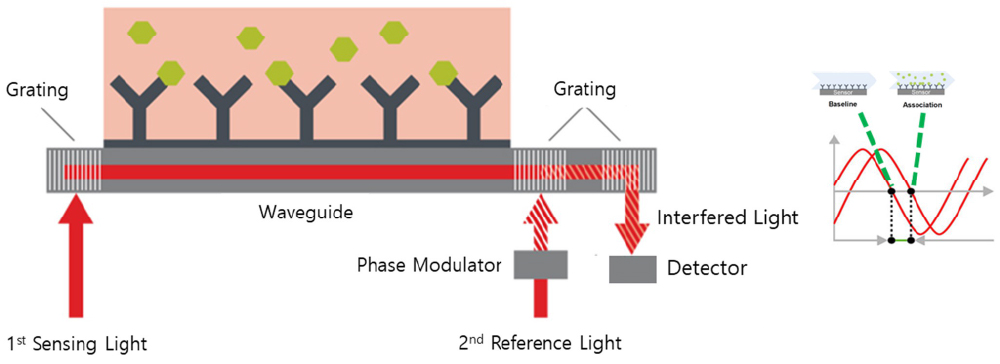
Grating Coupled Interferometry (GCI, 그레이팅 결합 간섭법) 원리 설명

- Sensing area (1mm2)의 넓은 범위에서 신호를 얻은 후, Reference light와 간섭을 일으켜 위상의 변화를 측정합니다.
- 만약 센서 표면의 물질 상호 작용이 일어나면, 위상의 변화(y축)를 시간(x축)에 따른 신호로 나타낼 수 있습니다.
- 높은 신호와 reference light와 간섭을 통해 high signal to Low noise ratio 및 높은 해상능을 실현했습니다 (noise < 0.01 pg/mm2, Rmax < 1 pg/mm2 (1 pg/mm2= 1RU))
- 본 WAVE system에는 고감도 성능의 GCI 센서를 탑재하여 보다 신뢰성이 높은 카이네틱스 분석이 가능합니다.
Label-free, Real-time Molecular Interaction analysis이란?
Label-free Sample?
형광을 붙인 probe나 HRP등을 붙인 antibody등의 라벨을 붙이지 않은 것을 의미합니다. 종전에는 샘플 각각에 대해 tagging을 필수적으로 이행해야 했습니다. 본 제품에서는 샘플 전처리 (Labeling)하지 않고 특이성을 가지는 물질에 대한 결합 반응을 측정할 수 있습니다.
Real-time Molecular Interaction analysis?
- 사용자의 분석 목적에 따라 약물 후보간 물질 결합 반응을 선별 (screening) 또는 특성 분석(characterization)의 목적으로 사용 가능합니다.
- 1개의 target 작용기 단백질 (Ligand라고 부름)을 chip에 고정시킨 뒤, 다수의 약물 후보군 (analyte라고 부름)를 순차적으로 흘려주어 결합 친화도를 분석할 수 있습니다.
- 작용기 단백질과 약물 후보간 결합 (association), 결합 평형상태 (equilibrium-state), 해리 (dissociation) 등의 결합 상태를 실시간으로 확인할 수 있습니다.
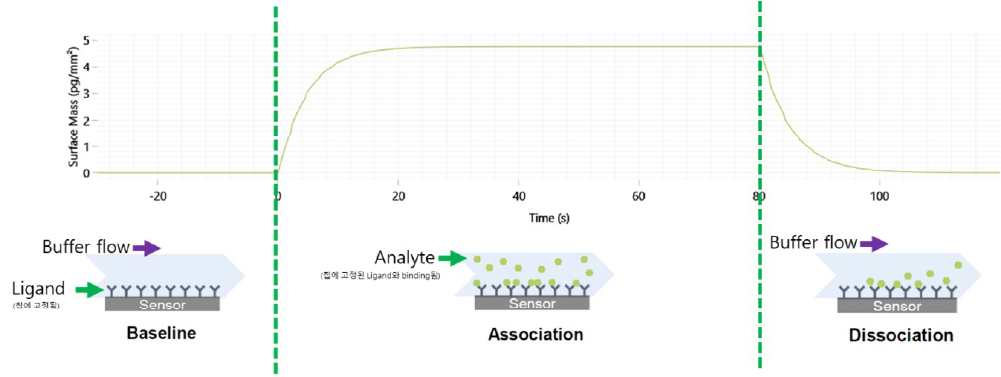
물질 결합 반응에 따른 실시간 Sensorgram 결과
사용예시
예를 들어 코로나 바이러스 치료제 개발하기 위해 작용기 단백질 (spike protein) 반응하는 물질을 찾는 연구를 한다고 가정해보겠습니다. 실험자는 WAVEchip에 작용기 단백질 (spike protein, ligand)을 고정시켜줍니다. 그리고 난 후에, 다수의 약물 후보군 (analyte)인, 다수의 천연물, 다수의 chemical compound, 다수의 Protein 후보 물질 등을 순차적으로 흘려주었습니다. 이 중 다수의 반응하는 물질을 1차 선별한 뒤, 후속 실험을 통해 최종 1개 물질을 치료제 후보로 선정할 수 있습니다. 이러한 예시를 통해 결합 반응하는 물질을 실시간으로 선별할 수 있으며, 특히, 얼마나 빠르게 붙는지, 늦게 떨어지는지 등의 affinity, kinetics 분석을 수행하여 약물의 필요 dose, duration time 등을 prediction할 수 있습니다.
고속 측정을 위한 waveRAPID® 분석법
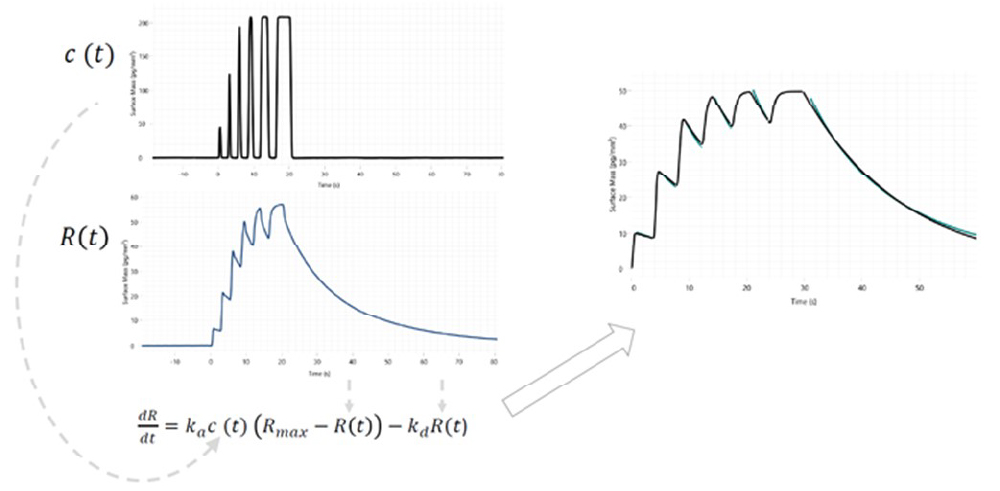
- WAVEdelta 모델에서만 가능한 waveRAID®방법은 기존 multi-cycle traditional kinetics (MCK)와 다른 획기적인 방법입니다.
- 1개 샘플당 1개 희석액 (higher starting concentration 1개)만 준비하면 됩니다.
- 1개 샘플을 sample loop로 빨아들인 후, 자동으로 pulse 시간을 조절하여 내뿜으로써 1회 중 다농도 분석을 실시하고, dissociation은 1회 수행하는 획기적인 방법입니다.
- 1 싸이클만으로 full kinetics (ka, kd, KD, Rmax) 분석이 가능한 획기적인 분석방법입니다.
- screening 중 MCK방법으로 수행 시, 1샘플 1농도로 수행하게되며, 이경우 full kinetics data를 얻지 못하고 단지 affinity 확인만 가능한 단점을 보완한 독특한 방법입니다.
- Creoptix사의 특허 방법입니다.
Regeneration-free kinetics 실험과 마찬가지로, 후속 분석 펄스가 도입되기 전에 응답은 기준선으로 돌아갈 필요가 없습니다. 단일 농도 주입 시간은 WAVEcontrol 소프트웨어에 의해 자동으로 조정됩니다. 이를 통해 단계 희석이 필요없고 1 웰에서의 카이네틱스 측정이 가능합니다.
셋업 시간을 대폭 단축하고,샘플 1개당 분석을 가속화하여, 보다 더 많은 샘플에 대해 분석 가능수 있습니다.
소프트웨어 WAVEcontrol®
WAVEcontrol은 유연성과 기능성을 결합한 워크플로 기반 소프트웨어입니다. 분석 설정, 데이터 평가부터 보고서 작성에 이르기까지 모든 단계를 직관적인 조작으로 쉽게 수행할 수 있습니다. 다음 기능이 포함됩니다.
- - 내장된 마법사 (wizard) 및 이전 실험에서 설정한 사이클 (preset)을 활용하고 사용자 지정하여 신속하게 분석 설정 가능
- - kinetics fitting을 위한 미리 정의된 모델 (1:1 interaction, mass transport, heterogenous ligand, conformational change, bivalent) 및 user customized evaluation model 사용 가능
- - Traditional fitting (Global fitting)으로 데이터 평가 또는 Direct Kinetics 도구를 사용한 자동 분석 가능
- - Excel 및 기타 형식으로 데이터 가져오기 및 내보내기
- - Open XML 형식의 파일을 사용하여 기존 LIMS에 데이터 통합 (Gene data Screener와 호환 가능)
- - 자동 보고 기능으로 사용자 지정 가능한 PDF 또는 Word 형식으로 결과 내보내기
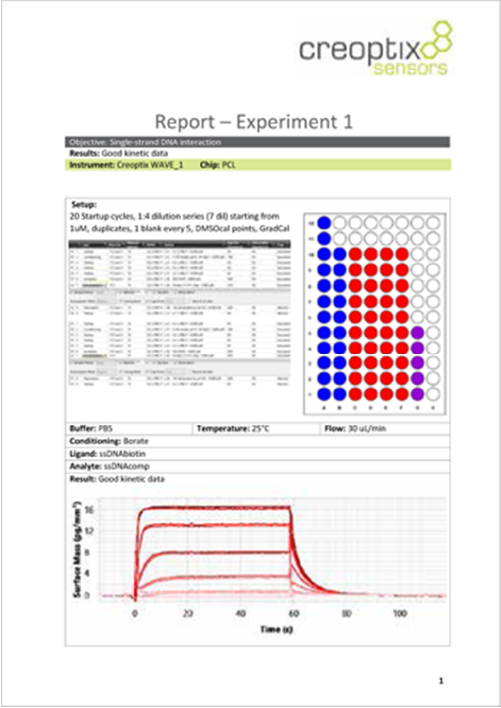
References
Small Molecules/FBDD
Nature Communications, 2021
C. Hong, NJ Byrne, B. Zamlynny, S. Tummala, L. Xiao, JM Shipman, AT Partridge, C. Minnick, MJ Breslin, MT Rudd, et al. “Structures of active -state orexin receptor 2 rationalize peptide and small-molecule agonist recognition and receptor activation.” DOI: 10.1038/s41467-021-21087-6
Small Molecules/FBDD
RSC Advances, 2021
EA Fitz Cederfelt, M. Abramsson, H. Ludviksdottir, JE van Muijlwijk-Koezen, IJP de Esch, D. Dobritzsch, T. Young, H. Danielson. “Discovery of fragments inducing conformational effects in dynamic proteins using a second-harmon .” DOI: 10.1039/D0RA09844B
Small Molecules/FBDD
Scientific Reports, 2020
H. Jankovics, B. Kovacs, A. Saftics, T. Gerecsei, E. Tóth, I. Szekacs, F. Vonderviszt, R. Horvath. “Grating-coupled interferometry reveals binding kinetics and affinities of Ni ions to genetically engineered protein layers.” DOI : 10.1038/s41598-020-79226-w
Plant Sciences, Peptides
PNAS, 2020
AD Crook, AC Willoughby, O. Hazak, S. Okuda, KR VanD NM Clark, R. Sozzani, M. Hothorn, CS Hardtke, ZL Nimchuk. “BAM1/2 receptor kinase signaling drives CLE peptide-mediated formative cell divisions in Arabidopsis roots.” DOI : 10.1073/pnas.2018565117
COVID-19 Research
bioRxiv, 2020
JD Walter, CAJ Hutter, AA Garaeva, M. Scherer, I. Zimmermann, M. Wyss, J. Rheinberger, Y. Ruedin, JC Earp, P. Egloff, M. Sorgenfrei, L. Hürlimann, I. Gonda, G. Meier, S. Remm, S. Thavarasah, G. Zimmer, DJ Slotboom, C. Paulino, P. Plattet, MA Seeger. “Highly potent bispecific sybodies neutralize SARS-CoV-2.” DOI : 10.1101/2020.11.10.376822
Biologics Development, COVID-19 Research
bioRxiv, 2020
JD Walter, CAJ Hutter, I. Zimmermann, M. Wyss, P. Egloff, M. Sorgenfrei, LM Hürlimann, I. Gonda, G. Meier, S. Remm, S. Thavarasah , P. Plattet, MA Seeger. “Sybodies targeting the SARS-CoV-2 receptor-binding domain.” DOI : 10.1101/2020.04.16.045419
Protein-Protein Interactions
Analyst, 2019
B. Peter, A. Saftics, B. Kovacs, S. Kurunczi, R. Horvath. “Oxidation increases the binding of EGCG to serum albumin revealed by kinetic data from label-free optical biosensor with reference channel.” DOI : 10.1037/ C9AN0
Plant Sciences, Peptides
The EMBO Journal, 2019
K. Lau, R. Podolec, R. Chappuis, R. Ulm, M. Hothorn. “Plant photoreceptors and their signaling components compete for binding to the ubiquitin ligase COP1 using their VP peptide motifs.” DOI: 10.15252/embj.2019102140
Biologics Development
Journal of Molecular Biology, 2019
F. Andres, L. Iamele, T. Meyer, JC Stüber, F. Kast, E. Gherardi, HH Niemann, A. Plückthun. “Inhibition of the MET Kinase Activity and Cell Growth in MET-Addicted Cancer Cells by Bi-Paratopic Linking.” DOI: 10.1016/j.jmb.2019.03.024
Plant Sciences
Nature Plants, 2018
U. Hohmann, J. Nicolet, A. Moretti, LA Hothorn, M. Hothorn, “The SERK3 elongated allele defines BIR ectodomains in brassinosteroid signalling.” DOI: 10.1038/s41477-018-0150-9
Small Molecules/FBDD
E-Life, 2018
W. Pitsawong, V. Buosi, R. Otten, RV Agafon Kern, S. Kutter, G. Kern, RAP Pádua, X. Meniche, D. Kern. “Dynamics of human protein kinase Aurora A linked to drug selectivity.” DOI: 10.7554/eLife.36656
※이 외에도 다수의 논문이 있습니다. 자세한 내용은 문의하시기 바랍니다.
Specification
Creoptix WAVE
General
- Noise (RMS)
- <0.01 pg/mm2 @ 1 Hz
- Drift
- <0.3 pg/mm2/min
- Readout Frequency
- 1 Hz, 10 Hz or 40 Hz
- Association Const. Range
- ka = 103 – 5x107 M-1 s–1 (small molecules)
ka = 103 – 3x109 M-1 s–1 (large molecules)
- Dissociation Const. Range
- kd = 10–5 – 10 s–1
- Analysis Temperature Range
- 15°C – 40°C
- Molecular Weight Limit
- No lower limit
- waveRAPID® Functionality
- No
Fluidics
- Flow Channels / Path
- 2, parallel
- Channel Referencing
- 1-4 and 4-1 or 2-3 and 3-2
- Flow Cells
- Sealed, disposable, integrated into disposable WAVEchip
- Flow Rate
- 1 – 400 µl/min
- Crude Sample Robustness
- Yes
Sample Handling
- Sample Capacity
- 2x microtiter plates
(96 or 384 well, standard or deep well) or vial racks (48 positions of 1.5ml)
- Buffer
- 1 buffer
- Degasser
- Built-in
- Injection Volume
- < 450 µl, 100 µl typical
- Sample Volume Required
- Injection volume plus 15-50 µl (application dependent)
- Sample Storage Temperature
- Ambient or 4°C – 20°C regulated
- Sample Recovery
- Yes
- Automation
- 120h of unattended operation
Data Treatment
- Information Provided
- Kinetic and affnity data (ka, kd, KD)
- Graphs
- Real-time curves, multiple curve overlays, fit, report point plots
- Data Extraction
- Curves, ka, kd, KD tables, graphs, reports
- Data Analysis
- Fully automated data evaluation
- Kinetic Models
- Predefined models including 1:1 interaction, mass transport, heterogenous ligand, conformational change and bivalent
- Direct Kinetics
- Yes
Creoptix WAVEdelta
General
- Noise (RMS)
- <0.01 pg/mm2 @ 1 Hz
- Drift
- <0.3 pg/mm2/min
- Readout Frequency
- 1 Hz, 10 Hz or 40 Hz
- Association Const. Range
- ka = 103 – 5x107 M-1 s–1 (small molecules)
ka = 103 – 3x109 M-1 s–1 (large molecules)
- Dissociation Const. Range
- kd = 10–5 – 10 s–1
- Analysis Temperature Range
- 4°C – 45°C (max 20°C below ambient)
- Molecular Weight Limit
- No lower limit
- waveRAPID® Functionality
- Yes
Fluidics
- Flow Channels / Path
- 4, parallel
- Channel Referencing
- Any combination of the 4 channels
- Flow Cells
- Sealed, disposable, integrated into disposable WAVEchip
- Flow Rate
- 1 – 400 µl/min
- Crude Sample Robustness
- Yes
Sample Handling
- Sample Capacity
- 2x microtiter plates
(96 or 384 well, standard or deep well) or vial racks (48 positions of 1.5ml)
- Buffer
- Automatic switching between 4 buffers
- Degasser
- Built-in
- Injection Volume
- < 450 µl, 100 µl typical
- Sample Volume Required
- Injection volume plus 15-50 µl (application dependent)
- Sample Storage Temperature
- Ambient or 4°C – 20°C regulated
- Sample Recovery
- Yes
- Automation
- 120h of unattended operation
Data Treatment
- Information Provided
- Kinetic and affnity data (ka, kd, KD)
- Graphs
- Real-time curves, multiple curve overlays, fit, report point plots
- Data Extraction
- Curves, ka, kd, KD tables, graphs, reports
- Data Analysis
- Fully automated data evaluation
- Kinetic Models
- Predefined models including 1:1 interaction, mass transport, heterogenous ligand, conformational change and bivalent
- Direct Kinetics
- Yes
Product Information
| Cat No. | 품명 | 규격 | 제조사 | 제품정보 |
|---|---|---|---|---|
| WAVE-2h | WAVEsystem | - | Creoptix |
Label-free, Real-time, Automated - Molecular Interaction Analyzer - Surface-based Microfluidic pump system - 2 Channel - GCI biosensor equipped - No-Clog Microfluidics consumables (WAVEchip) - 1-Click operation and evaluation software: WAVEcontrol |
| WAVEdelta-4ch | WAVEdelta | - | Creoptix |
Label-free, Real-time, Automated - Molecular Interaction Analyzer - Surface-based Microfluidic pump system - 4 Channel - GCI biosensor equipped - No-Clog Microfluidics consumables (WAVEchip) - 1-Click operation and evaluation software: WAVEcontrol - waveRAPID analysis Included (Prepare 1 dilute per 1 sample, Inject once, and Obtain 1 full kinetic data set) |
Releated Product
WAVEchip® 라인업
다양한 샘플을 이용한 어플리케이션을 위해 다양한 표면의 센서 칩을 준비하고 있습니다.
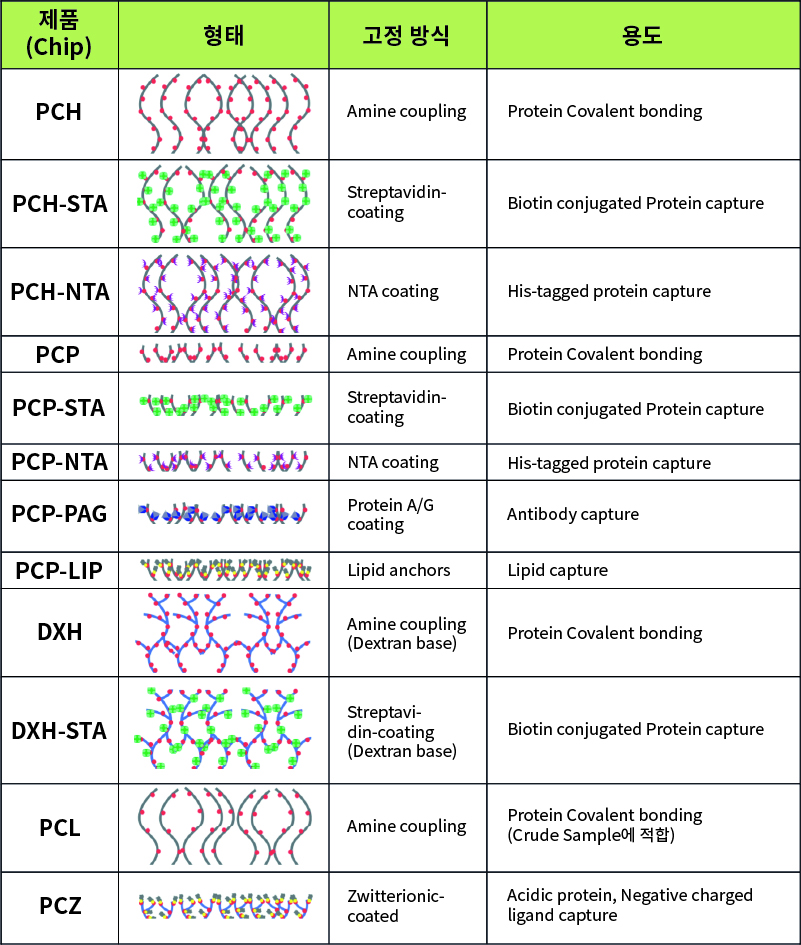
PC*: Polycarboxylate 재질
DX*: Dextran 재질